Share

Talking Techniques
How can we achieve gender equality in STEM?
This International Women’s Day takeover episode, with special guest host BioTechniques’ Senior Digital Editor Abi Sawyer, takes a look at the results of Future Science Group’s (London, UK) survey for the scientific community on gender equality and parity in STEM.
Abi’s guests on this episode are the Vice President of Epidemiology and Clinical Evidence at IQVIA (NC, USA), Dr Christina Mack; the Executive Director for the Pharmaceutical Research Computing Center at the University of Maryland School of Pharmacy (MD, USA), Dr Ebere Onukwugha; a Lecturer, Science Communicator and Author based in Cardiff (UK), Dr Emma Yhnell; and the Director of the Neuroscience Center Microscopy Core at the University of North Carolina–Chapel Hill (NC, USA), Dr Michelle Itano.
They discuss the results of Future Science Group’s survey, share their own experiences of gender inequality as well as situations where they’ve felt supported, and outline how the STEM community can push further towards gender equality and parity.
Contents:
Introduction: 00:00 – 1:47
Gender inequality, experiences and impact on STEM careers: 1:48 – 10:41
Underlying reasons for gender disparity in STEM: 10:42 – 17:51
Promoting gender equality in the STEM work environment: 17:52 – 30:34
Looking forward: representation and mentoring: 30:35 – 39:32
Conclusions: 39:33 – 40:10
More episodes
View all episodes

Cytokine networks in autoimmune diseases: mechanisms, pathogenesis and therapeutic innovations
21:54|In this episode of Talking Techniques, Ritwika Biswas, Field Application Scientist at Sino Biological US Inc. (PA, USA), discusses the role of cytokines in autoimmune diseases, the techniques used to examine them and some emerging therapeutic innovations beginning to change the way we approach the treatment of autoimmune diseases.ContentsIntroduction: 00:00–02:06The role of cytokines in a healthy body: 02:06–03:57Cytokines in autoimmune diseases: 03:57–06:24Techniques for detecting cytokines in autoimmune diseases: 06:24–09:48Targeting cytokines for therapeutic purposes: 09:48–11:54Challenges with targeting cytokines in autoimmune diseases: 11:54–14:28Addressing the challenges of targeting cytokines: 14:28–16:43Established cytokine-targeting drugs: 16:43–18:57The future of cytokines in autoimmune diseases: 18:57–21:54
Skills-based teaching and microcredentialing in STEM
54:22|This episode of Talking Technique deviates slightly from specific lab technologies to instead discuss techniques and methods we use for teaching and testing life sciences.To do this, I’m speaking to two pioneers of unconventional teaching and testing approaches to STEM education. Angela Consani is the Co-Founder and CEO of the Bioscience Core Skills Institute (KS, USA). This skills-first microcredential program provides certification for lab skills in techniques, safety and quality control, using performance-based practical testing. Natalie Kuldell is the Founder and Executive Director of Biobuilder (MA, USA), a nonprofit organization, set up to increase interest, understanding and engagement in STEM by converting lab research projects in into teachable modules aimed primarily at the pre-graduate level to give students the practical skills needed for a career in the life sciences.Together, we’ll question the current system of STEM education and training and whether it captures all the potential talent that could be channeled into the life sciences, best serving all the roles available in the industry.Contents:Introductions: 00:00-03:00Introducing BioBuilder: 03:00-07:00What industry wants from skills-based testing: 07:00-11:25How well do current university degrees meet these requirements: 11:25-15:40Designing curriculums to meet the requirements of industry and updating life science education to meet the demands of a new world: 15:40-21:55The practicalities of a skills-based curriculum: 21:45-23:50Conducting skill-based testing: 23:50-28:40Testing BioBuilder’s curriculum: 28:40-32:00Can skills-based courses really provide the underlying knowledge needed to flourish in a career in STEM: 32:00-37:00How the biotech industry is responding to skills-based teaching and testing: 37:00-46:00The interplay between testing and learning and industry: 46:00-51:20Outro: 51:20-54:00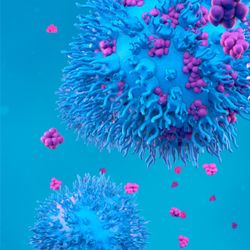
6. Antigen validation and T-cell receptor engineering for cancer immunotherapies
19:41||Season 5, Ep. 6This episode of the Talking Techniques podcast dives into the realm of cancer immunotherapies, focusing on antigen discovery and T-cell receptor engineering for T-cell therapies. Guiding us through the field is Jim Heath, President of the Institute for Systems Biology in Seattle, where he runs the Heath Lab, investigating fundamental immunology, and infectious and chronic diseases. Jim discusses the computational models and wet lab techniques he uses to characterize T cells, the importance of targeting a balanced immune response with immunotherapies and more in this podcast recorded at AACR 2024 (5th–10th April 2024; San Diego, CA, USA).Contents:Introductions: 00:00-02:00Intro to cancer vaccines and T-cell therapies: 02:00-04:00Antigen detection and validation in T-cell therapies: 04:00-05:20Wet lab and computational techniques for antigen detection: 05:20-09:15The importance of a balanced immune response to cancer immunotherapies: 09:15-10:30Technological developments in antigen detection: 10:30-13:45 Tips for best practice when conducting T-cell receptor design 13:45-15:40What is one thing you would like to see change in the field of antigen detection and T-cell receptor engineering? 15:40-16:30 Designing the path towards a more balanced immune response from immunotherapies 16:30-19:40
4. Cytokines: from therapeutics to diagnostics
26:11||Season 5, Ep. 4In this episode of Talking Techniques, Ritwika Biswas, Field Application Scientist at Sino Biological US Inc. (PA, USA), discusses the use of cytokines in immunotherapy. Ritwika details the role of cytokines in the body, before going on to discuss how they can be used as therapeutics and to guide treatment decisions. Ritwika also shares how she thinks these proteins will be used in the future.Contents· Introduction: 00:00–01:35· The role of cytokines in the body: 01:35–02:52· Immune regulation and signaling: 02:52–05:40· Cytokine interactions and networks: 05:40–08:42· Modulating cytokine activity for therapeutic purposes: 08:42–12:35· The influence of cytokines on immunotherapy outcomes: 12:35–16:04· Using cytokines to predict treatment responses and guide immunotherapy decisions: 16:04–20:44· The importance of standardizing and validating cytokine diagnostic assays: 20:44–24:36· The future of cytokines in immunotherapy: 24:36–26:11
3. Spatial analysis of the immune-cell-surface proteome at a single-cell resolution
23:58||Season 5, Ep. 3The cell-surface proteome plays a critical role in immune-cell function; however, our ability to examine its interactions and spatial organization has previously been limited by available proteomic techniques. This episode explores the function of immune-cell membrane proteins and how the latest developments in spatial proteomics have enabled more detailed interrogation of these proteins and their spatial relationships.Our guest, Hanna van Ooijen, Immunology Application Scientist at Pixelgen Technologies guides us through the field, revealing a new technique that enables spatial analysis of the cell-surface proteome at a single-cell resolution and highlighting some exciting discoveries that it has facilitated.Contents:Introductions: 00:00-01:40Introducing Molecular Pixelation: 01:40-02:15Example applications of Molecular Pixelation: 02:15-03:20The role of membrane proteins in immune cell function: 03:20-07:25Traditional techniques to investigate cell membrane proteins: 07:15-10:20Recent improvements in investigative technology and our understanding of immunology: 10:20-11:10Challenges associated with current technologies: 11:10-13:50How Molecular Pixelation can address these challenges: 13:50-15:25Molecular Pixelation workflow: 15:25-17:55Tips for best practice when using molecular pixelation: 17:55-19:30Exciting discoveries using Molecular pixelations: 19:30-21:00Potential implications of molecular pixelation for the future of immunology: 21:00-24:00
2. Investigating the neurological pathways underlying vocal communication
34:27||Season 5, Ep. 2In this episode of Talking Techniques, we catch up with Michael Long, Principle Investigator of the Long Lab at New York University (NY, USA), where he investigates the neural circuits that underlie vocal communication.Through the examination of animal models, from songbirds to the rare singing mice of Costa Rica, with cutting-edge imaging techniques Michael reveals fascinating insights into vocal communication. We also discuss his human experiments, working alongside neurosurgeons, with emerging electrophysiological probes to monitor the neural activity of participants as they speak and interact, ultimately revealing how this research could begin to provide solutions for neurological conditions impacting communication, such as autism.Contents:Introduction: 00:00 – 01:40Investigating neural circuits underlying vocal communication: 01:40 – 04:15Techniques to explore animal models of vocal communication: 04:15 – 06:25The impact of cooling brain regions on songbird singing: 06:25 – 07:50The techniques used to investigate animal models: 07:50 – 12:20Songbirds: 07:50 – 09:45The singing mouse: 10:00 – 12:20Investigating neural circuits in humans during speech: 12:20 – 16:30Investigating neural circuits in humans during conversation: 16:30 – 19:00Moving beyond neural area identification towards understanding neural pathways and mechanisms: 19:00 – 21:40Navigating neuropixels, big data and safety: 21:40 – 26:10If there was one thing you could ask for to help you better understand these pathways, what would it be? 26:10 – 27:55The experience of working with patients undergoing neurosurgery: 27:55 – 30:30The potential impact on speech disorders and autism: 30:30 – 33:15
1. Rare disease and pharmacogenomics
20:18||Season 5, Ep. 1Launching our fourth season of Talking Techniques, this episode, supported by the University of Cincinnati (OH, USA) we delve into rare disease research and pharmacogenomics, their intersection and the key techniques used to explore them.Guiding us through these fields is Brenna Carey, an Assistant Professor at Cincinnati Children’s Hospital Medical Center whose research focuses on rare disease pathogenesis, diagnostics and therapeutic development and who also runs key courses on the University’s Pharmacogenomics and Drug Discovery Masters degree programs.Contents:Introduction: 00:00-01:15An introduction to pulmonary alveolar proteinosis (PAP) and rare lung diseases 01:15-03:50Techniques to investigate the pathogenesis of PAP: 03:50-05:30Developing diagnostics and therapeutics for PAP: 05:30-08:20The importance of pharmacogenomics in drug development: 08:20-11:25Key techniques and approaches in pharmacogenomics: 11:25-13:00Emerging trends in pharmacogenomics: 13:00-15:05Key takeaways from your pharmacogenomics course: 15:00-18:00What would you ask for to improve our understanding of pharmacogenomics? 18:00-20:15This episode is supported by the University of Cincinnati Online
9. One man’s waste in another man’s treasure: using wastewater to monitor infectious diseases
20:47||Season 4, Ep. 9In this episode of Talking Techniques, we talk to Andrew Lee, a senior research fellow in Queen’s University Belfast’s (UK) wastewater-based epidemiology group, about his work using wastewater to monitor and detect infectious diseases. Andrew discusses how wastewater surveillance acts as an early warning system, providing novel, unbiased insights into human and animal pathogens that are circulating within a community, and how this can contribute to a ‘One Health’ approach. He also explains how he has incorporated nanopore sequencing into his work, and the advantages that this provides.Contents:· 00:00–01:45: Introductions· 01:45–03:45: Wastewater surveillance for infectious disease· 03:45–05:35: Genomic surveillance approaches can complement established epidemiological methods· 05:35–07:25: Why look at wastewater?· 07:25–10:40: The advantages of nanopore sequencing for wastewater surveillance· 10:40–12:25: The experimental workflow· 12:25–15:05: Using wastewater surveillance to detect both human and avian influenza· 15:05–18:20: Wastewater surveillance as an early warning system· 18:20–20:47: Future perspectives: other environmental samples, antimicrobial resistance and what else can be found in wastewater?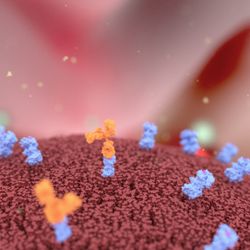
8. Next-generation antibody therapeutics
24:52||Season 3, Ep. 8In this episode of Talking Techniques, we speak to two experts from Sino Biological US Inc. (PA, USA) about the latest developments in antibody technologies and how these developments have led to the next generation of antibodies that are revolutionizing therapeutic approaches to a number of diseases.With the guidance of Field Scientist Ritwika Biswas and Technical Account Manager Grace Liu, we explore the challenges of developing and working with next-generation antibodies, the latest developments and applications of these molecules and the holy grail that antibody designers are driving towards.Contents:Introduction: 00:00 – 02:40The history of monoclonal antibody therapeutics: 02:40 – 04:40The working principles of multi-specific antibodies: 04:40 – 08:15Recent developments in ADCs: 08:15 – 11:35Challenges with the development of multi-specific antibodies and ADCs: 11:35 – 13:55Solutions to address these challenges: 13:55 – 16:25Clinical applications of multi-specific antibodies and ADCs: 16:25 – 20:30The dream of real-time adaptability for the next generation of antibody therapeutics: 20:30 – 24:52